Understanding and managing your class list is important for maintaining consistent annotations.
Why Class Indices Matter (ifcb-analysis Users)
Note: This section is primarily relevant if you export
.matfiles for use with the ifcb-analysis MATLAB toolbox (Sosik & Olson, 2007). If you use the default SQLite storage or work with CSV exports, class indices are less critical because class names are stored directly.
IFCB .mat annotations use numerical indices to reference classes:
Index 1 → unclassified
Index 2 → Ciliate
Index 3 → Dinoflagellate
Index 4 → Diatom
...When you save an annotation as a .mat file for ifcb-analysis, the file stores:
- ROI number: 00042
- Class index: 2 (meaning “Ciliate”)
If you change the order or remove classes, existing .mat annotations become invalid for ifcb-analysis!
Creating a Class List from Scratch
You can create a new class list directly in the app without uploading a file:
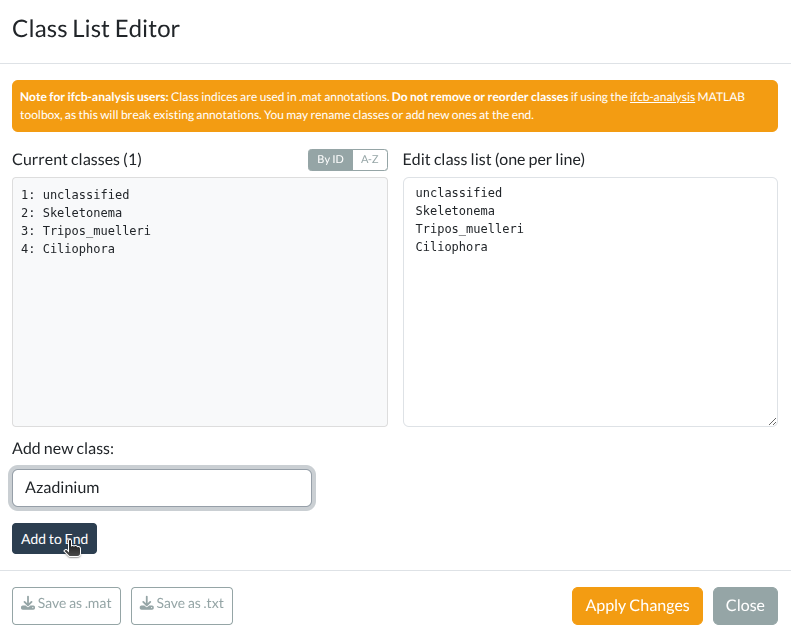
Class list editor showing classes with annotation counts, editing area, and export options. Click to enlarge.
- Open Settings → Edit Class List
- The editor opens with an empty class list
- Add classes using either method:
Method 1 - One at a time:
- Type a class name in the “Add new class” field
- Click Add to End
- Repeat for each class
Method 2 - Bulk entry:
- Type or paste class names in the text area (one per line)
- Click Apply Changes
- The app creates a temporary class list file automatically
- If using SQLite storage (default), your class list is auto-saved to the database and restored on next startup
Note: With SQLite storage, your class list persists automatically. You can still use Save as .mat or Save as .txt to export a portable copy for sharing or backup.
Loading an Existing Class List
Viewing Classes
- Open Settings → Edit Class List
- The left panel shows all classes with their indices and the number of annotated images per class (queried from the database)
- Toggle By ID / A-Z to sort the view
Note: Sorting is for viewing only - it doesn’t change actual indices.
Editing Classes
Renaming a Class
You can safely rename classes without breaking annotations:
- Find the class in the edit text area
- Change the name
- Click Apply Changes
The index stays the same, so existing annotations remain valid.
Adding New Classes
New classes must go at the end of the list:
Method 1 - Quick Add:
- Type name in “Add new class” field
- Click Add to End
Method 2 - Text Edit:
- Add new classes at the bottom of the text area
- Click Apply Changes
What NOT to Do (ifcb-analysis Users)
Note: These restrictions apply if you export .mat files for use with ifcb-analysis. CSV exports store class names directly and are not affected by index changes.
- Never remove classes - This shifts all subsequent indices
- Never reorder classes - This changes index-to-name mapping
- Never insert in the middle - This shifts indices below
Example: What Goes Wrong (ifcb-analysis)
Original list:
1: unclassified
2: Ciliate
3: Dinoflagellate
4: DiatomIf you remove “Ciliate”:
1: unclassified
2: Dinoflagellate ← Was index 3, now index 2!
3: Diatom ← Was index 4, now index 3!Now all your “Ciliate” (index 2) .mat annotations become
“Dinoflagellate” when loaded in ifcb-analysis!
WoRMS Matching (AphiaID)
You can match class names to the World Register of Marine Species directly in the Class List Editor:
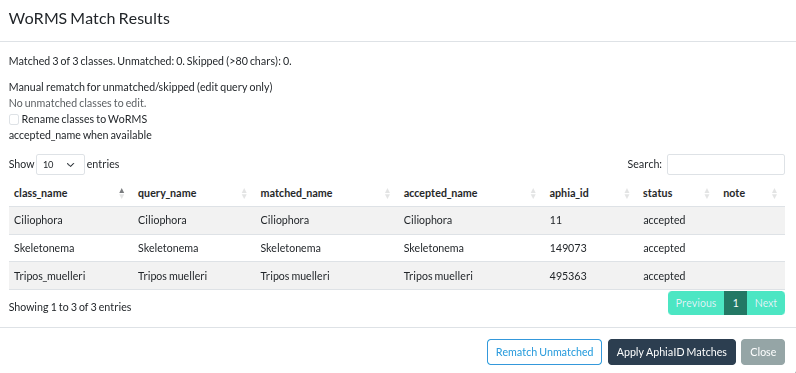
WoRMS match results modal with manual rematch query fields for unresolved classes. Click to enlarge.
- Open Settings → Edit Class List
- Click Match WoRMS AphiaID
- Review the result table:
-
accepted: direct accepted WoRMS match -
synonym: class matched through a synonym; accepted AphiaID is returned -
unmatched: no WoRMS match found -
skipped: query skipped automatically (e.g. name longer than 80 characters)
-
- For unresolved rows, edit the query text in the manual rematch fields and click Rematch Unmatched
- Click Apply AphiaID Matches
After applying, matched classes show [AphiaID: ...] in
the class list display.
Where this is stored: AphiaID mappings are saved in
the SQLite database (class_taxonomy table), not in
class2use.mat/.txt.
Exporting Class Lists
Save as MAT
Creates MATLAB-compatible class2use.mat for use with ifcb-analysis:
- Click Save as .mat
- Choose location in browser download
Note: Saving .mat files requires Python with scipy.
Auto-persistence in SQLite
When the storage format includes SQLite (the default), the class list
is automatically saved to the global_class_list table in
annotations.sqlite whenever it changes. This means:
- Adding a class, renaming, applying WoRMS matches, or uploading a file all persist instantly
- On next startup, the class list is restored from the database (no file upload needed)
- File-based class lists (
.mat/.txt) are still supported as a fallback and for sharing
If you switch away from SQLite storage, class list persistence falls
back to the file-based workflow (upload or save as
.txt/.mat).
Best Practices
Backup your class list before making changes
Add classes at the end only (required for ifcb-analysis compatibility)
Never delete - mark deprecated classes with prefix like “OLD_” (for ifcb-analysis users)
Document changes in your project notes
Use consistent naming - decide on convention (underscores, capitalization)
Tip: If you don’t use ifcb-analysis and only work with CSV exports, you have more flexibility with class list management since CSV files store class names directly.
Class Naming Conventions
Allowed Characters
Most characters commonly used in taxonomic names are allowed:
- Letters (a-z, A-Z)
- Numbers (0-9)
- Underscores (
_) - Hyphens (
-) - e.g., “Strombidium-like” - Periods (
.) - Spaces
Characters That Are Modified
Since class names are used as folder names when exporting PNGs, some characters are automatically sanitized:
| Character | Why | What Happens |
|---|---|---|
/ or \
|
Path separators would create subdirectories | Replaced with _
|
< > " ' & |
HTML/security risks | Removed |
: * ? \| |
Invalid on Windows | Removed |
.. |
Path traversal security risk | Removed |
Example for ambiguous taxa: If you have
Snowella/Woronichinia in your class list, it will be saved
as Snowella_Woronichinia. The app will display a message
when this happens.